VCF Explorer
I built a tool to bridge the gap between computational methods and and biologists investigating genomic variants.
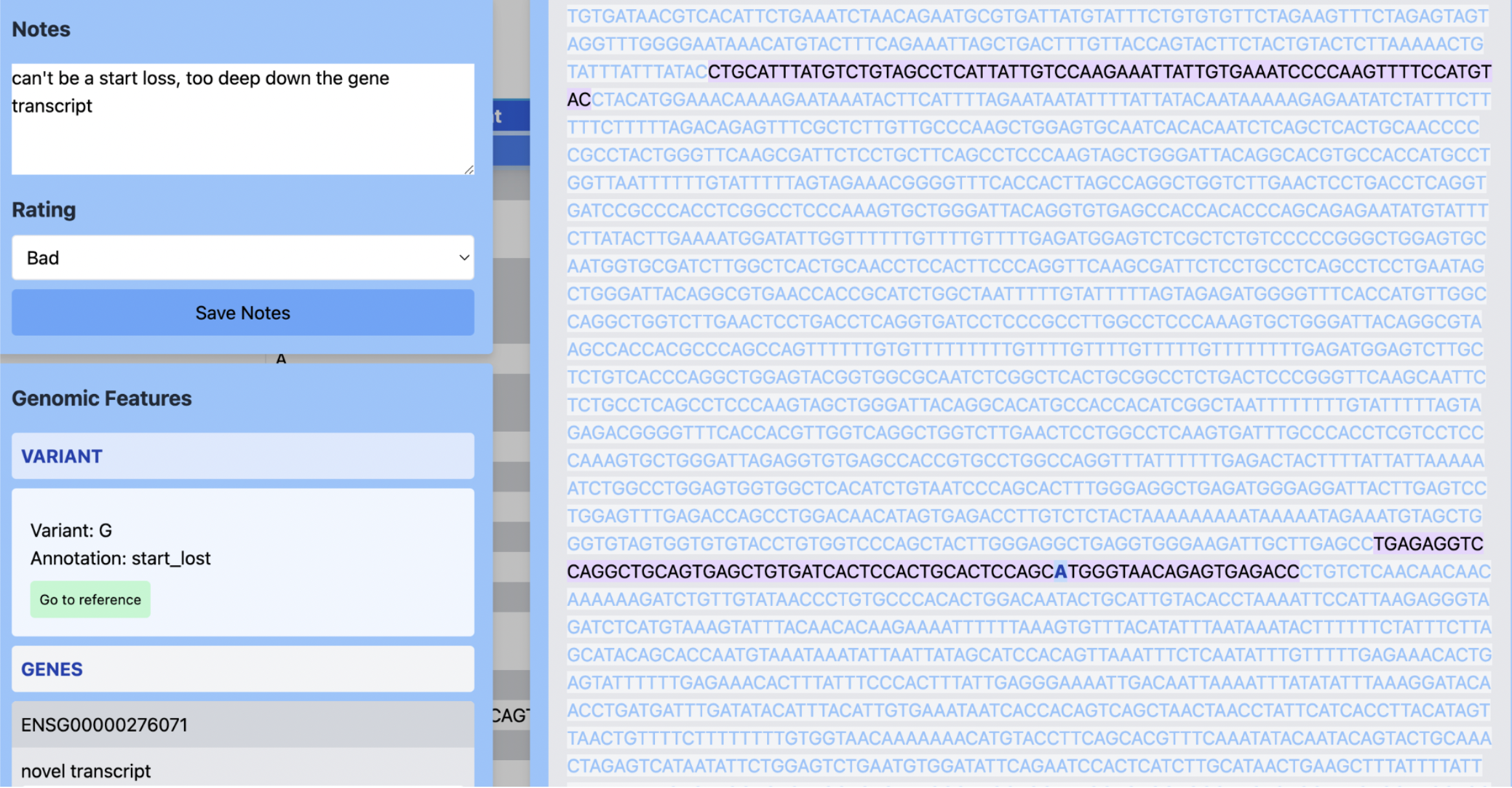
The tool is an alternative to command line tools or traditional genome explorers, which often look like this:
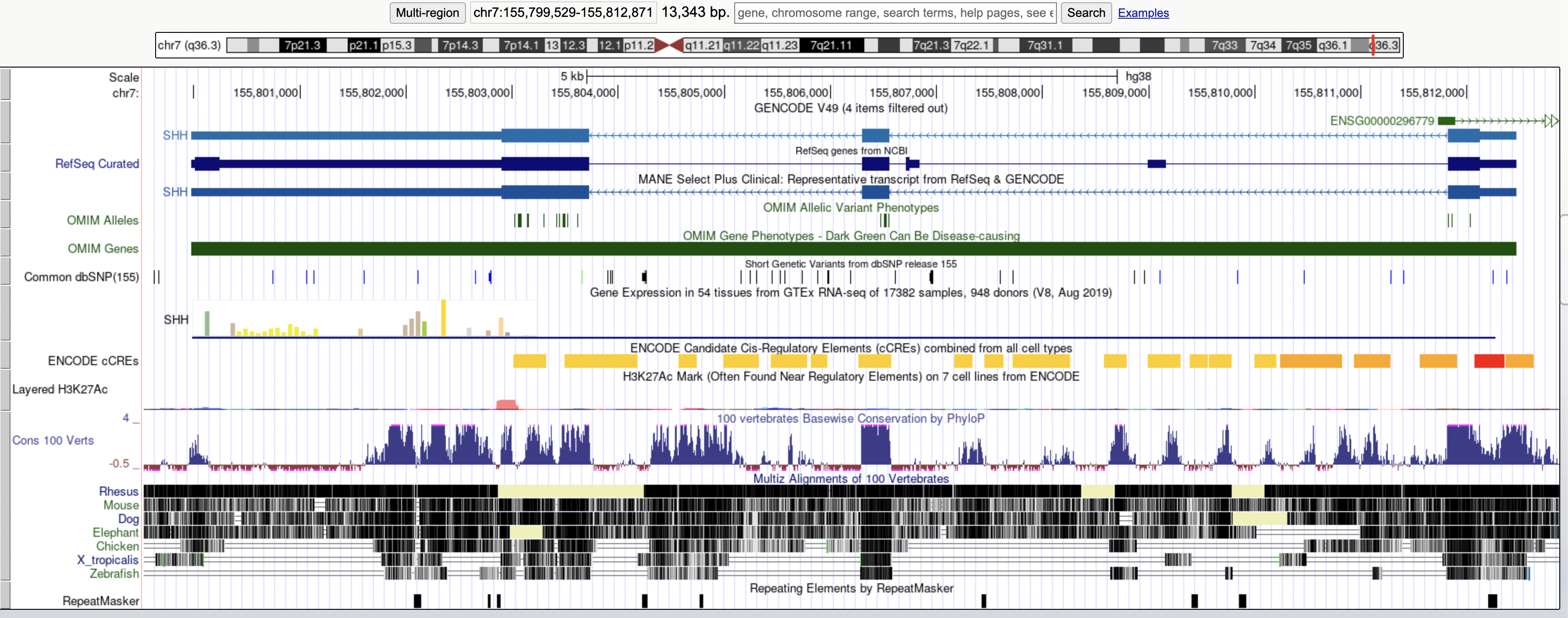
It makes investigating the impact of variants more user friendly and intuitive by:
-
fast loading and navigation to the context around each variant position
-
nucleotide-level and codon-level visibility. The variant can be seen within the larger genomic context, including the sequence for the whole gene, annotations, transcripts, alternative transcripts, coding regions
-
built-in rating and note-taking tool to record and share notes on each variant
-
removing visual noise